A groundbreaking international research collaboration has revealed an unexpected molecular mechanism employed by an obscure lineage of terrestrial plants, offering a potentially transformative pathway for genetically re-engineering staple food crops like rice and wheat to dramatically enhance their photosynthetic efficiency and, consequently, their yield. This discovery centers on a novel approach to optimizing Rubisco, the planet’s most critical carbon-fixing enzyme, which has long been a bottleneck in agricultural output.
The Imperative for Photosynthetic Enhancement
Global food security remains one of the most pressing challenges of the 21st century. With a burgeoning global population projected to reach nearly 10 billion by 2050, coupled with the escalating impacts of climate change, the demand for increased agricultural productivity is paramount. Traditional methods of yield improvement, such as expanding arable land or intensifying fertilizer use, are increasingly unsustainable or reaching their practical limits. Consequently, scientists are exploring fundamental biological processes to unlock new avenues for crop enhancement. Among these, improving photosynthesis – the process by which plants convert sunlight, water, and carbon dioxide into biomass – stands out as a frontier with immense potential. Even a modest increase in photosynthetic efficiency could translate into substantial gains in crop yields, reducing the pressure on natural resources and mitigating environmental footprints.
At the heart of photosynthesis lies Rubisco (ribulose-1,5-bisphosphate carboxylase/oxygenase), an enzyme universally recognized as the primary entry point for nearly all organic carbon into the global food web. Despite its pivotal role in sustaining life on Earth, Rubisco is famously inefficient. It operates at a relatively slow pace and, crucially, exhibits a significant affinity for oxygen in addition to carbon dioxide. When Rubisco binds with oxygen, it initiates a wasteful process known as photorespiration, which consumes energy and previously fixed carbon compounds, thereby diminishing the plant’s overall photosynthetic output and growth rate. This inherent flaw in Rubisco represents a major limitation to the productivity of C3 plants, which include most of the world’s major food crops like wheat, rice, and soybeans.
Evolutionary Solutions and Engineering Challenges
Nature, in its vast evolutionary laboratory, has devised several strategies to mitigate Rubisco’s shortcomings. For instance, C4 plants, such as maize and sugarcane, and CAM plants (Crassulacean Acid Metabolism), found in cacti and succulents, have evolved complex biochemical and anatomical adaptations to concentrate carbon dioxide around Rubisco, effectively reducing photorespiration. Another elegant solution is observed in many types of algae and cyanobacteria, which sequester Rubisco within specialized subcellular compartments called pyrenoids. These microscopic structures function as carbon-concentrating mechanisms (CCMs), actively pumping carbon dioxide into the pyrenoid, thereby creating a high-CO2 environment that favors Rubisco’s carboxylation activity over its oxygenation activity.
For decades, researchers have harbored ambitions of transferring these naturally evolved CCMs into C3 food crops. The goal has been to engineer a more efficient Rubisco environment within crop plant cells, mimicking the success seen in C4 plants or algae. However, attempts to transplant the intricate molecular machinery of algal pyrenoids, which often involve dozens of genes and complex protein interactions, into land plants have proven exceptionally challenging. The sheer complexity of these systems and the vast evolutionary distance between algae and higher land plants present formidable genetic engineering hurdles. The intricate interplay of proteins, transporters, and structural components required for a functional pyrenoid has largely defied direct transfer, necessitating a search for simpler, more evolutionarily compatible mechanisms.
Hornworts: A Surprising Evolutionary Bridge
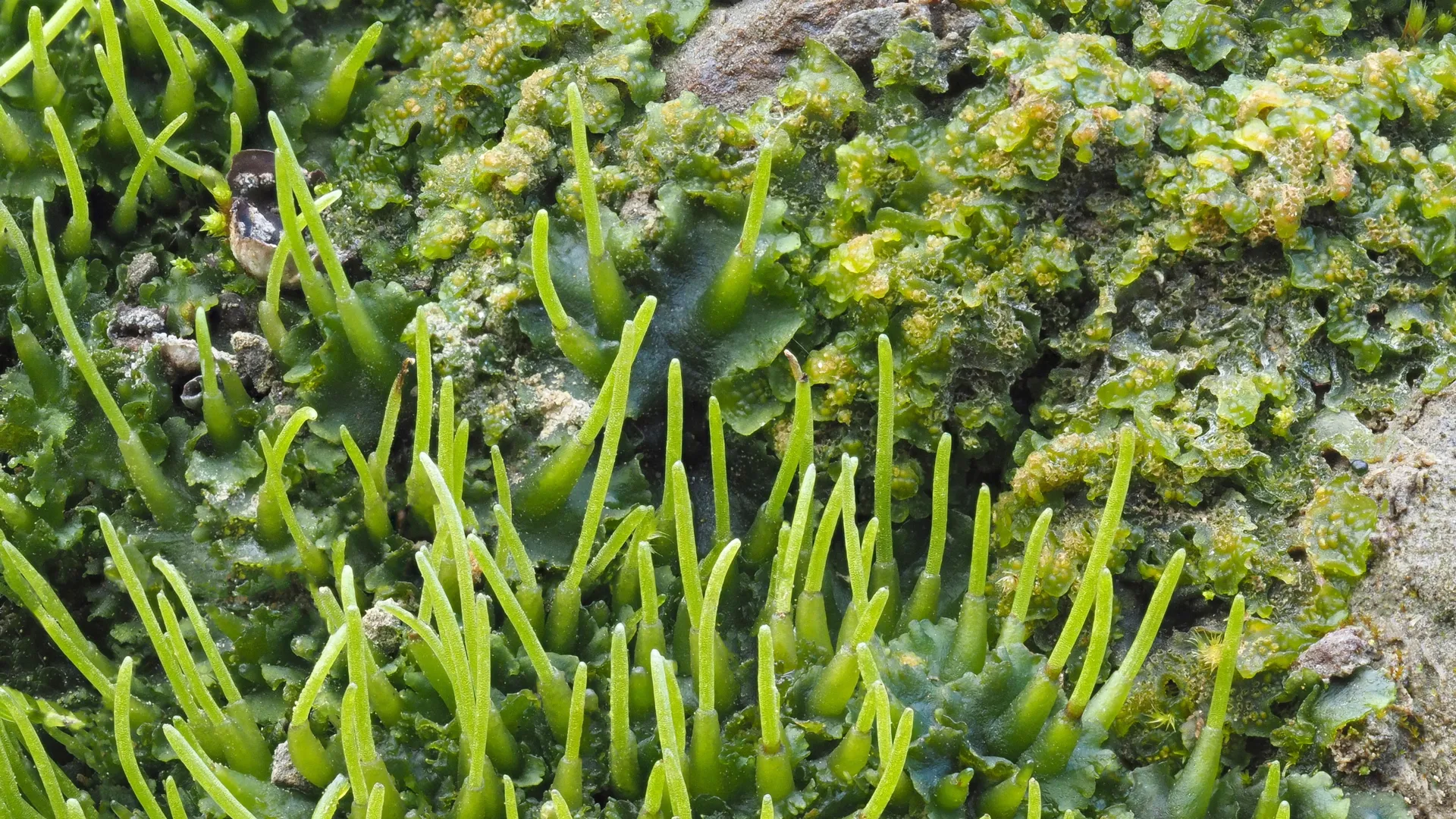
The recent breakthrough emerged from an unexpected corner of the plant kingdom: hornworts. These ancient, inconspicuous bryophytes are unique among land plants in possessing subcellular structures that bear a striking resemblance to the pyrenoids found in algae. This observation immediately captured the attention of the scientific community because hornworts, despite their primitive appearance, share a much closer evolutionary lineage with modern crop plants than algae do. This phylogenetic proximity suggested that any carbon-concentrating strategies employed by hornworts might be mechanistically simpler and, critically, more amenable to genetic transfer into agricultural species.
Initial hypotheses posited that hornworts would employ a mechanism analogous to their algal counterparts, utilizing a separate, dedicated scaffold protein to aggregate Rubisco into these concentrated compartments. However, the international research team, comprising scientists from the Boyce Thompson Institute (BTI), Cornell University, and the University of Edinburgh, discovered a profoundly different and surprisingly elegant strategy. Instead of relying on an external scaffolding protein, hornworts have intrinsically modified Rubisco itself to facilitate its clustering.
The RbcS-STAR Mechanism: Molecular Velcro in Action
The crux of this discovery lies in an unusual protein component identified by the researchers and named RbcS-STAR. Rubisco is a complex enzyme typically composed of multiple large and small protein subunits. In hornworts, a specific variant of the small subunit (RbcS) incorporates an additional, unique C-terminal segment, which the team designated the STAR (Small subunit Tail-Associated Rubisco) region.
This STAR region functions as a kind of molecular velcro. Its presence causes individual Rubisco holoenzymes to physically associate and self-assemble into larger, dense clusters within the plant cell’s chloroplasts. This intrinsic clustering mechanism is a radical departure from the externally templated assembly observed in algal pyrenoids, where a separate, non-Rubisco protein acts as the central organizing principle. The direct modification of a Rubisco subunit to confer self-assembly capabilities represents a highly parsimonious and potentially more tractable evolutionary solution.
To rigorously validate the function and modularity of the RbcS-STAR component, the research team conducted a series of critical experiments. First, they introduced the RbcS-STAR component into a closely related hornwort species that naturally lacks pyrenoids. The result was striking: Rubisco, which was previously dispersed throughout the chloroplasts, reorganized itself into distinct, concentrated structures visually analogous to natural pyrenoids. This experiment provided compelling evidence that the STAR region alone was sufficient to induce Rubisco clustering in a relevant biological context.
Further expanding their investigation, the scientists tested the RbcS-STAR mechanism in Arabidopsis thaliana, a widely utilized model plant in laboratory research, phylogenetically much closer to crop plants than hornworts. Remarkably, when the hornwort RbcS-STAR component was expressed in Arabidopsis, the native Rubisco proteins within the chloroplasts also aggregated into dense, localized compartments. This finding underscored the modularity and potential universality of the STAR region. In an even more refined experiment, researchers demonstrated that merely attaching the isolated STAR tail to Arabidopsis‘s native Rubisco small subunit was enough to trigger the same clustering effect. This definitively established the STAR region as the independent, driving force behind Rubisco aggregation, confirming its potential as a plug-and-play molecular tool.
Towards a New Generation of High-Yield Crops
The discovery that the RbcS-STAR mechanism functions effectively across different plant species, from hornworts to Arabidopsis, holds profound implications for agricultural biotechnology. It suggests that scientists may be able to induce Rubisco clustering in major crop plants by simply introducing this relatively small, modular protein component. This approach bypasses the immense complexity of transferring entire algal pyrenoid systems, offering a more direct and potentially faster route to engineering improved photosynthetic efficiency.
However, the researchers are careful to emphasize that Rubisco clustering is but one piece of the puzzle in constructing a fully functional carbon-concentrating mechanism. While the STAR region provides the "house" for Rubisco, the efficiency of this "house" will depend on its ability to maintain a high concentration of carbon dioxide around the clustered enzyme. As aptly described by one of the co-leaders of the research, Laura Gunn of Cornell University, "We have built a Rubisco house, but it won’t be an efficient house unless we update the HVAC." This analogy highlights the crucial next steps: developing robust mechanisms for actively delivering and concentrating carbon dioxide within these newly formed Rubisco clusters. This will likely involve engineering specialized CO2 transporters or carbonic anhydrase enzymes that can rapidly convert bicarbonate to CO2 and sequester it effectively within the clustered environment.
A Sustainable Future for Food Production
Despite the remaining challenges, the identification of the RbcS-STAR mechanism represents a monumental leap forward in the quest to enhance photosynthesis. Even a marginal increase in the efficiency with which staple crops convert sunlight into biomass could have cascading benefits. Higher yields from existing agricultural lands could reduce the need for further deforestation, preserving biodiversity and natural ecosystems. Enhanced efficiency could also lead to reduced inputs of water and fertilizers per unit of food produced, thereby lowering the environmental footprint of agriculture. Furthermore, crops with more robust photosynthetic machinery might exhibit greater resilience to abiotic stresses, such as heat and drought, which are becoming increasingly prevalent due to climate change.
This research exemplifies the power of investigating the often-overlooked corners of the natural world for innovative solutions. Nature has, through millions of years of evolution, already experimented with and refined countless biochemical strategies. The role of scientific inquiry is to meticulously uncover these strategies, understand their underlying principles, and then judiciously apply that knowledge to address humanity’s most pressing challenges. By leveraging the elegant simplicity of the hornwort’s molecular velcro, scientists are now closer than ever to engineering a new generation of super-efficient crops, laying a critical foundation for a more sustainable and food-secure future for all. The comprehensive study detailing these findings was recently published in the esteemed journal Science, underscoring its significance within the scientific community.